Open Data in Science
Presentation slides from Make.OpenData.ch camp 2012
Information overload in science
Modern science produces enormous amounts of data. Only in Life Sciences 2500-3000 new research articles are published every day (see PubMed articles published in the last year). These articles are in the plain-text pdf format, and hence can only be interpreted by humans, which makes knowledge dissemination rather slow.
Non-transparency of academic research
An average project in academia takes 2-5 years before publication of results. During this period research process is non-transparent to people outside of the research group. Because of this, several groups may be doing exactly the same work without knowing about duplication of their efforts. Also, because negative results are almost never published – scientists in different groups would [uselessly] repeat same experiments, despite these have already been tried by others.
<html> Information overload and non-transparency of research process makes scientific innovation increasingly inefficient.</html>
Possible remedy
One could solve the above issues if scientists would start openly sharing ALL their data, in ~real-time and in machine-readable (semantically annotated) form. Then, intelligent algorithms/tools could be developed which would automatically find and recommend scientists information which is relevant to their research. This would minimize duplication of research efforts and reduce the information overload.
Showcasing “Open Science + Linked Data” to science policy makers
Science funding agencies are increasingly recognizing the problems of non-transparency and inefficiency of academic research. Open Access to research articles (a subset of the global Open Science vision) is already being demanded by many science funding agencies (1, 2). Unfortunately there are almost no examples of how “real-time” Open Science combined with Semantic technologies could enhance the efficiency of scientific innovation.
<html> The goal of proposed project for make.OpenData.ch is to create a basic demo of such “Open Science + Linked Data” combination. </html>
Future developments
The vision behind this “Open Science + Linked Data” demo is supported by a number of academic researchers, including group leaders, in the field of Structural Biology. We hope to evolve this vision towards a sustainable activity in the future (e.g. development of intelligent information management tools for [academic] science).
Data
- Currently - scraping of local data + linking it to UniProt
- Experimental data used for the demo comes from a research project in the field of Structural Biology performed at Uni Basel and ETH Zurich. If successful the code of this demo will be deployed to provide continuous web “publishing” + visualization of research data obtained in a specific research project in Structural/Systems Biology at ETH Zurich.
Possible goals for the make.OpenData.ch hackday
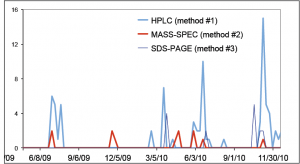
- make a time-course visualization of data generated by several different methods in the course of specific research project (e.g. image below).
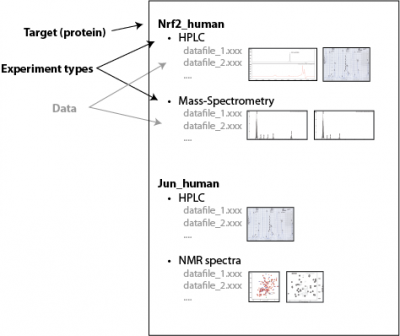
- make a structured view of the above data – grouping by specific experiments and biological targets (e.g. proteins, see image below).
- annotate the data (methods and proteins) using namespaces of existing bio-ontologies (Gene Ontology and/or Ontology of Biomedical Investigations).
- provide links from the above structured data to related data from Life Science databases in the Linked Open Data cloud (e.g. UniProt, PubMed).
Possible technologies for demo (from a lame point of view 
- script parsing filenames in a given folder (server or local) to extract experiment / object identities > e.g. Python.
- html representation of extracted data + annotation using RDFa and namespaces of GO and/or OBI ontologies
- visualization of performed experiments vs time > e.g. dynamic graph with Javascript and HTML5
- perl / python / ruby querying of bio databases & establishment/visualization of links between local data objects and related information in public databases.